The Gut Microbiome and Early-Onset Colorectal Cancer: Mechanisms, Biomarkers, and Clinical Implications
by Hannah Zhong1#, Jianan Li2#, Kyung-il Kim3, Pranati Shah4, Claire Jung5, Lucas Kim6, Dani Castillo7*, Gagandeep Brar7*
1City of Hope, Department of Hematology and Hematopoietic Cell Transplantation, 1500 E Duarte Rd, Duarte, CA 91010, USA.
2University of Toronto, Faculty of Kinesiology & Physical Education, 100 Devonshire Place, Toronto, Ontario, CA.
3Kaiser Permanente Fontana Medical Center, Department of Internal Medicine, 9961 Sierra Ave, Fontana, CA 92335, USA.
4Loma Linda University Medical Center, Department of Internal Medicine, 11234 Anderson St, Loma Linda, CA 92354, USA.
5Flintridge Preparatory School, 4543 Crown Ave, La Cañada Flintridge, CA 91011, USA.
6Loma Linda University, School of Medicine, 11175 Campus St, Loma Linda, CA 9235, USA.
7City of Hope, Department of Medical Oncology and Therapeutics Research, 1500 E Duarte Rd, Duarte, CA 91010, USA.
#These authors contributed equally to this work.
*Corresponding author: Brar G, City of Hope, Department of Medical Oncology and Therapeutics Research, 1500 E Duarte Rd, Duarte, CA 91010, USA.
Received Date: 24 February, 2026
Accepted Date: 04 March, 2026
Published Date: 06 March, 2026
Citation: Zhong H, Li J, Kim KI, Shah P, Jung C, et al. (2026) The Gut Microbiome and Early-Onset Colorectal Cancer: Mechanisms, Biomarkers, and Clinical Implications. J Oncol Res Ther 11: 10332. DOI: https://doi.org/10.29011/2574-710X.10332
Abstract
Early onset colorectal cancer (EOCRC), now accounting for a rapidly increasing proportion of global CRC incidence, remains a major clinical concern due to delayed diagnosis, aggressive tumor biology, and limited screening strategies in younger adults. Increasing evidence demonstrates a strong association between the gut microbiome dysbiosis and EOCRC initiation and progression. This review highlights the role of microbial composition, microbial metabolites, and host-microbe interactions as emerging biomarkers with diagnostic and therapeutic relevance in EOCRC. Overall, integrating microbiome profiling into early detection, risk stratification, and targeted prevention may improve outcomes in EOCRC.
Implications for Practice
EOCRC is rising in incidence and frequently diagnosed at advanced stages, highlighting the limitations of current screening strategies. Growing evidence indicates that EOCRC is associated with distinct gut microbiome signatures that interact with host genetics and environmental exposures to promote tumorigenesis. These findings suggest that microbiome-informed approaches could enhance risk stratification, support earlier detection, and refine prevention strategies in younger adults. Although microbiome-based interventions are not yet ready for routine clinical use, the integration of microbial profiling may ultimately inform precision screening and therapeutic decision making in EOCRC.
Keywords: Colorectal Cancer; Early Onset Colorectal Cancer; Gut Microbiome; Biomarkers; Dysbiosis, Metabolites;
Introduction
Colorectal cancer (CRC) is the third most common cause of cancer-related death worldwide, with an estimated 1.9 million new cases and over 930,000 deaths annually [1]. While overall CRC incidence in older adults has declined, cases among younger individuals have risen sharply, with early-onset CRC (EOCRC), defined as diagnosis at age 50 or younger, now accounting for up to 12% of new CRC diagnoses. The incidence is increasing, making it the second leading cause of cancer death in individuals under 50 by 2030 [2]. Patients with EOCRC often present at more advanced stages due to initial nonspecific symptoms and delayed recognition [3]. Because of this, outcomes for many patients with EOCRC remain poor, and treatment is often complicated by long-term survivorship concerns, fertility issues, and unique tumor biology compared to later-onset CRC (LOCRC).
The objective of this review is to evaluate the current evidence linking the gut microbiome to EOCRC, with a focus on microbial species, metabolites, and dysbiosis patterns that may drive tumor initiation and progression. We will also highlight emerging data on microbiome-based biomarkers and therapeutic strategies, emphasizing their potential to improve risk prediction, prevention, and treatment of EOCRC.
The Microbiome in Early Onset Colorectal Cancers
The gut microbiome is increasingly recognized as a key contributor to the pathogenesis of EOCRC [4]. In healthy individuals, the intestinal microbiota maintains epithelial barrier integrity, regulates host metabolism, and modulates both local and systemic immunity. This complex ecosystem supports homeostasis through short-chain fatty acid (SCFA) production, immune regulation, and protection against pathogenic overgrowth [5].
In EOCRC, disruption of this equilibrium, or dysbiosis, appears to play a particularly important role. Emerging evidence reveals that the microbial composition of EOCRC differs markedly from that of LOCRC, reflecting both taxonomic and functional divergence as summarized in Table 1. Evidence from sequencing and metagenomic studies suggests that the microbiome in EOCRC may differ in both composition and functional potential compared with late-onset CRC.
Younger patients often exhibit depletion of beneficial, SCFA-producing commensals alongside enrichment of inflammatory and genotoxic species. Fecal microbiome analyses demonstrate that younger patients harbor communities enriched in bacteria associated with epithelial barrier disruption, inflammation, and altered energy metabolism.
In contrast, LOCRC microbiomes are characterized by a gradual depletion of short-chain fatty acid-producing commensals that normally support epithelial repair and immune balance [10]. Studies have highlighted overrepresentation of Fusobacterium nucleatum, enterotoxigenic Bacteroides fragilis (ETBF), and Escherichia coli harboring pks islands in younger CRC patients, with concomitant depletion of butyrate-producing taxa such as Faecalibacterium prausnitzii and Roseburia [6, 7]. These microbial alterations are thought to contribute to tumorigenesis through chronic inflammation, genotoxic metabolite production, and disruption of epithelial barrier function [8, 9].
These compositional distinctions are also evident within tumor-associated microbiomes. A Chinese cohort identified Flavonifractor plautii as a distinguishing microbial marker of EOCRC and noted higher relative abundance of Streptococcus species in younger patients [11, 12]. EOCRC tumors also show greater alpha diversity than those from older patients, indicating colonization by a more taxonomically and functionally diverse microbial community. Many of these taxa are implicated in mucin degradation, energy metabolism, and immune signaling, suggesting a dynamic and interactive tumor-microbiome interface [13]. LOCRC tumors are often dominated by species linked to chronic inflammation and genotoxicity, reflecting the cumulative effects of long-term dysbiosis and aging [14].
|
Feature |
EOCRC |
LOCRC |
|
Age group |
≤50 years |
>50 years |
|
Microbial trends |
↑ Fusobacterium nucleatum, Bacteroides fragilis, Escherichia coli (pks+), |
↓ SCFA producers |
|
Akkermansia muciniphila, Flavonifractor plautii; ↓ Faecalibacterium prausnitzii, Roseburia, Bifidobacterium adolescentis |
||
|
Functional profile |
↑ Virulence genes, genotoxin production, inflammation; ↑ mucin degradation and barrier disruption |
↓ Butyrate synthesis, ↑ chronic inflammation |
|
Pathways affected |
Wnt/β-catenin, NF-κB, STAT3, DNA damage response |
Age-related DNA methylation drift, oxidative stress |
|
Tumor |
Greater microbial diversity; metabolic reprogramming; early immune evasion |
Lower diversity; accumulation of chronic insults |
|
features |
Table 1: Distinct Microbial and Molecular Landscape of EOCRC vs. LOCRC; EOCRC, early-onset colorectal cancer; LOCRC, late-onset colorectal cancer; NF-κB, nuclear factor kappa-light-chain-enhancer of activated B cells; pks+, polyketide synthase gene-positive; SCFA, short-chain fatty acid; STAT3, signal transducer and activator of transcription 3; Wnt, Wingless/Integrated signaling pathway.
Microbial Metabolites as Drivers of Early Tumorigenesis
Host genetic alterations play a critical role in shaping EOCRC development alongside disruptions in the gut microbiome and environmental exposures, as illustrated in Figure 1. These cancer-driving mutations interact with microbial and immune signaling networks, creating tumor-promoting feedback loops that appear particularly accelerated in younger patients.
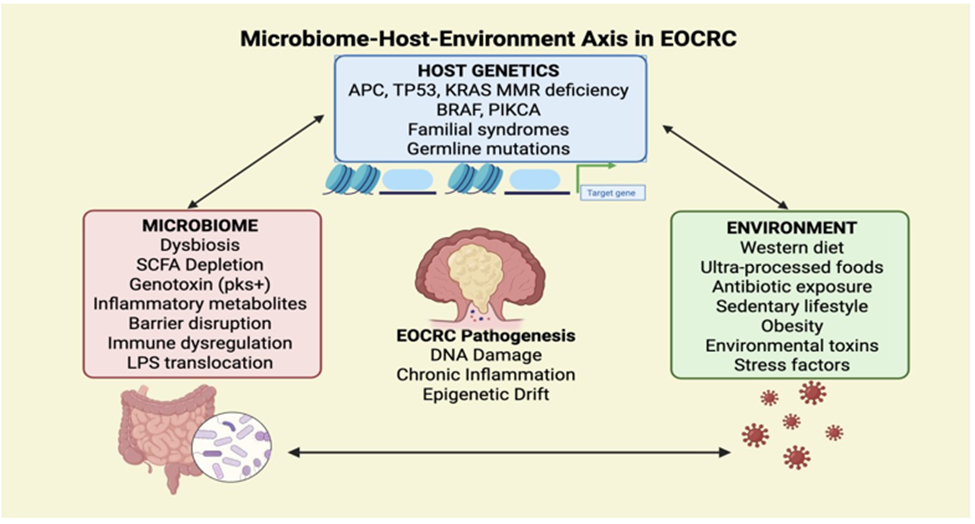
Figure 1: Microbiome-Host-Environment Axis in EOCRC.
Microbial metabolites arise from the bacterial breakdown of dietary and host-derived substrates, with major classes including SCFAs, polyamines, ethanol derivatives and bile acids [15, 16]. Disruptions in the balance of these metabolites can promote carcinogenesis by driving inflammation, oxidative stress, and epigenetic remodeling.
SCFAs, produced by fermentation of dietary fiber, normally maintain epithelial barrier integrity, regulate immune tolerance, and function as histone deacetylase (HDAC) inhibitors that suppress proliferative signaling [16]. In EOCRC, reduced abundance of SCFA-producing bacteria leads to lower butyrate availability, impairing apoptosis and promoting epithelial hyperproliferation. Reduced butyrate also alters chromatin accessibility and DNA repair capacity, accelerating early tumor initiation.
Microbial polyamines, including putrescine, spermidine, and spermine, support rapid cell turnover, but when produced excessively, enhance genomic instability and DNA alkylation [16]. Ethanol-derived acetaldehyde, another microbial metabolite, forms mutagenic DNA adducts and interferes with DNA repair processes, linking microbial ethanol metabolism to early colorectal tumorigenesis.
Microbial metabolism of primary bile acids into secondary bile acids, such as deoxycholic acid (DCA) and lithocholic acid (LCA), contributes to carcinogenesis through oxidative DNA damage and activation of nuclear receptors including FXR and TGR5 [17]. These metabolites promote cell proliferation, NF-κB-mediated inflammation, and epithelial stress signaling. Westernized microbiomes enriched in bile acid-metabolizing bacteria (e.g., Clostridium scindens) further amplify these effects, correlating with early epithelial transformation in younger patients. These metabolite-driven mechanisms are summarized in Figure 2, highlighting the interplay between protective and pathogenic microbial functions in EOCRC pathogenesis.
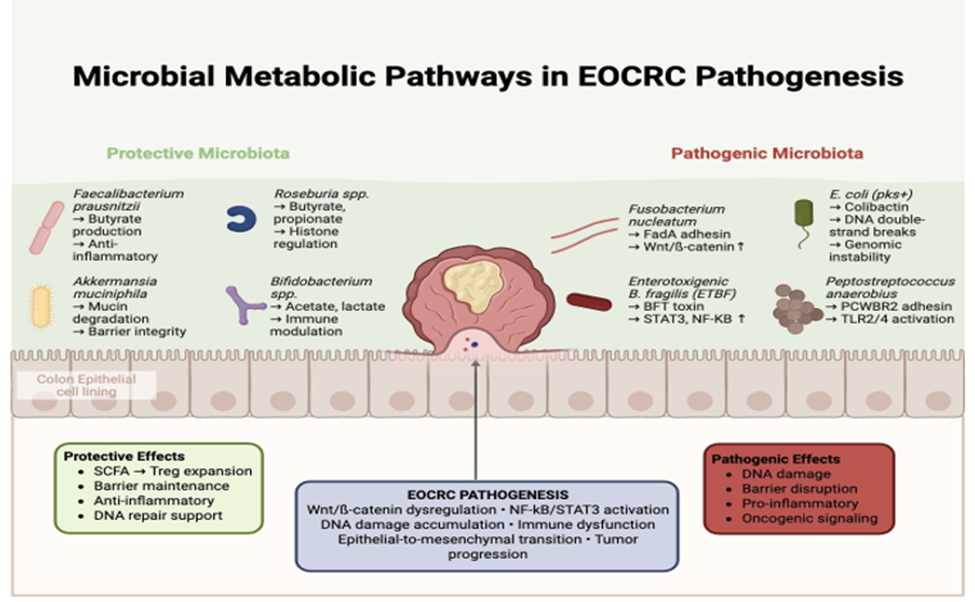
Figure 2: Microbial Metabolic Pathways in EOCRC Pathogenesis.
SCFA, short-chain fatty acid; Treg, regulatory T cell; NF-κB, nuclear factor kappa-light-chain-enhancer of activated B cells; STAT3, signal transducer and activator of transcription 3; BT toxin, Bacteroides fragilis toxin; TLR2/4, Toll-like receptor 2/4; FadA, Fusobacterium adhesin A; PCWBR2, Peptostreptococcus anaerobius cell wall-binding repeat protein 2.
Microbial Signatures of Risk Factors in EOCRC
Established risk factors include obesity, hereditary predisposition such as Lynch syndrome, inflammatory bowel disease, and lifestyle exposures including western diet, alcohol, and tobacco use.
Obesity
Obesity has repeatedly been associated with microbial changes that contribute to EOCRC. A study by Xu et al. suggested that EOCRC exhibits reduced fecal alpha diversity in the luminal gut microbiome compared with both LOCRC and healthy controls, with an increase in specific taxa that correlate with obesity and high-fat diet (HFD) [19]. Specifically, members of the Firmicutes phylum, including Blautia, Actinomyces, and Clostridia, showed positive correlations with HFD-induced obesity, while beneficial SCFA producing taxa were markedly reduced. The study also suggested that pathobionts relevant to EOCRC are responsive to obesity, such as Fusobacterium nucleatum, enterotoxigenic Bacteroides fragilis, and Escherichia coli, along with cohort signals indicating that Fusobacteria and Bacteroides differentiate EOCRC from LOCRC.
Notably, taxa associated with short-chain fatty acids (SCFAs) are diminished, and Mitsuokella multacida was not present in EOCRC samples Mitsuokella multacida was notably absent from EOCRC samples, further reflecting depletion of protective commensals.
Obesity-associated dysbiosis enriches pathobionts that activate Wnt/β-catenin and dampen anti-tumor immunity (Fusobacterium nucleatum), cleave E-cadherin and drive NF-κB/STAT3/IL-17 inflammation (enterotoxigenic Bacteroides fragilis), and deliver colibactin-mediated DNA damage (Escherichia coli pks⁺). These mechanisms provide a biological basis for why obesity may specifically accelerate EOCRC.
Lynch Syndrome
Lynch syndrome (LS), the most common hereditary cause of EOCRC, is characterized by germline deficiencies in DNA mismatch repair (dMMR) machinery. Studies in LS have revealed distinct microbial changes in patients with CRC. Increasing evidence suggests that dMMR-related genomic instability interacts with gut microbial dysbiosis to promote early neoplastic transformation.
In LS families, stool samples from CRC cases exhibit an increase in Agathobacter, Coprococcus, Ruminococcus, Lactobacillus, Fusobacterium, Parvimonas, and Peptostreptococcus, while showing a decrease in Bacteroides. Mouse models of LS additionally show that a combination of mismatch repair (MMR) deficiency and inflammation results in the expansion of Escherichia-Shigella, Akkermansia, Bacteroides, and Parabacteroides, whereas beneficial groups like Lactobacili and Lachnospiraceae are decreased [27]. In addition, Akkermansia muciniphila may degrade the mucus barrier under inflammatory conditions, exposing dMMR epithelium to microbial products and sustaining NF-κB–driven inflammation, while Parabacteroides contributes via succinate production and pro-inflammatory metabolic signaling, correlating with tumor burden in LS models.
While many studies focus on LS-associated CRC in general, various mechanisms explain why cancers in this group often arise earlier, contributing to EOCRC risk. MMR-deficient epithelium seems exceptionally reactive to microbial metabolites, with butyrate, usually considered as protective, serving as an oncometabolite that promotes hyperproliferation and accelerates neoplasia in LS mouse Models. Bacteria residing in crypts and the biofilm-like colonization in the proximal colon additionally enhance chronic inflammation, which, when combined with MMR deficiency, increases tumor burden. These findings match with mechanisms described in the mechanism section, including microbial activation of oncogenic signaling pathways. Therefore, in Lynch syndrome, defective dMMR fosters a vulnerable environment for early tumorigenesis. Collectively, these pathways suggest that although direct study of EOCRC in LS is lacking, the interaction between host genetics and microbial ecology may play a role in the increased risk and earlier development of CRC in this group.
Inflammatory Bowel Disease
Patients with inflammatory bowel disease (IBD) exhibit distinct changes in their gut microbiota that are believed to increase the risk of colorectal cancer. Because IBD frequently develops during adolescence or early adulthood, patients experience a longer cumulative period of dysbiosis and chronic inflammation, which may amplify cancer risk at young ages. In various cohorts, IBD correlates with reduced microbial diversity, a decline in beneficial butyrate-producing species like Faecalibacterium prausnitzii, Eubacterium Halli, and Roseburia, and an increased presence of pro-inflammatory bacteria like adherent-invasive Escherichia coli (AIEC) [36].
AIEC adhere to epithelial cells via type 1 pili binding to CEACAM6, invade epithelial cells, and live within macrophages, where they trigger TNF-a, IL-6, and IL-1 β secretion leading to chronic mucosal inflammation. Chronic inflammation from adherent-invasive Escherichia coli links to CRC by driving ROS/RNS-mediated DNA damage, cytokine-driven proliferation, and barrier dysfunction. However, limited studies have shown mechanisms that link AIEC to EOCRC. Additionally, spatial studies have shown that in IBD, bacteria are found closer to the epithelium and even within immune cell compartments, which enriches proteobacteria and colibactin-producing strains [37-39]. These microbial shifts may contribute to EOCRC via colibactin mutagenesis, barrier impairment, and chronic inflammatory signaling.
Diet/Lifestyle
Diets and lifestyle factors like Western dietary patterns, alcohol intake, and smoking are recognized to affect the gut microbiome. However, most existing studies investigate the relationship between these factors and CRC rather than EOCRC. Western diets are known for high fat and low fiber composition, associated with lower microbial diversity, fewer beneficial butyrate-producing taxa, and increased levels of pathobionts like Fusobacterium nucleatum and Enterococcus faecalis in CRC populations [40].
Enterococcus faecalis is known to play a role in colorectal carcinogenesis by producing reactive oxygen species that harm host epithelial DNA. A study by Wang et al showed that E. faecalis can trigger aneuploidy and tetraploidy in colonic epithelial cells via a bystander effect, where infected macrophages secrete extracellular superoxide and other reactive species that spread into adjacent epithelial cells. This oxidative stress causes chromosomal instability, breaks in DNA double strands, and disturbances in the mitotic process. Consequently, E. faecalis promotes a pro-tumorigenic setting marked by genomic instability, a crucial feature of colorectal cancer [41].
Dietary habits established during adolescence also shape long-term microbial and metabolic trajectories. High consumption of sugar-sweetened beverages has been associated with increased EOCRC incidence, accompanied by microbial shifts favoring saccharolytic, pro-inflammatory taxa and reduced SCFA production [42]. SSBs promote insulin resistance, increase oxidative stress, and enhance the presence of saccharolytic and pro-inflammatory taxa, while weakening butyrate-producing commensals.
In addition, a low intake of dietary fiber decreases SCFA production, impairs epithelial function, and removes a crucial anti-inflammatory mechanism for colonocyte [43]. This differs from LOCRC where years of unhealthy eating increase cancer risk, indicating that the microbiome-related effects of adolescent nutrition could specifically increase rate of EOCRC development.
Cigarette smoking and alcohol consumption have also been associated with higher levels of Bacteroides and reduced amounts of SCFA producing genera like Ruminococcaceae and Phascolarctobacterium, as well as lower fecal SCFA concentrations in healthy individuals [42]. These microbial changes are biologically important factors in tumorigenesis, yet no extensive studies have directly analyzed diet or lifestyle related microbial patterns in EOCRC patients, resulting in a significant knowledge gap. Proposed pathway links these exposures to carcinogenesis, including DNA damage from ethanol metabolism, barrier disruption, and SCFA depletion.
Antibiotics
The use of antibiotics is among the most significant environmental influences on the gut microbiome. Long term or repeated antibiotic use in childhood and adolescence has been associated with higher risks of colorectal adenomas and EOCRC [44, 45]. Antibiotics act by decreasing microbial diversity, inhibiting protective SCFA generating taxa, and promoting the proliferation of pathobionts like Fusobacterium nucleatum or toxigenic Bacteroides fragilis. Importantly, antibiotic effects occur more significantly in early life than later. Individuals with non-secretor FUT2 variants, who exhibit altered mucosal glycosylation and different microbiota composition, show stronger connections between antibiotic consumption and EOCRC risk.
Molecular Signatures of EOCRC
TP53
TP53 mutations are common in EOCRC, occurring in approximately 79.5% of EOCRC cases. These mutations lead to the accumulation of a dysfunctional p53 protein, impairing its role in cell cycle regulation and apoptosis. Studies have identified specific microbial taxa associated with TP53 mutations such as Eubacterium and Actinomyces species. These microbes may modulate local immune responses and contribute to epithelial integrity, potentially interacting with p53-driven tumor suppression pathways [46, 47].
Actinomyces can adhere to and invade mucosal surfaces, altering epithelial barrier function and promoting chronic inflammation. In a TP53-deficient setting, where apoptosis and DNA repair are compromised, this persistent inflammation enhances oxidative stress and allows damaged cells to survive and proliferate [48].
Eubacterium coprostanoligenes is a cholesterol-metabolizing bacterium that converts cholesterol into coprostanol, thereby influencing bile acid composition. Altered bile acid pools can modulate signaling through farnesoid X receptor (FXR) and TGR5 pathways which regulate immune responses and epithelial proliferation. Loss of p53-mediated control over inflammatory and metabolic signaling may amplify the effects of these microbial metabolites, fostering a pro-tumorigenic microenvironment [49].
PIK3CA
PIK3CA mutations are observed commonly in EOCRC. These mutations activate the PI3K/AKT signaling pathway, promoting cell proliferation and survival [50]. It has been found that compared to wild-type cases, PIK3CA mutated EOCRC has higher abundances of Megasphaera and Lactobacillus, while Actinomyces is reduced. These microbial taxa can influence PI3K/AKT signaling indirectly via metabolic or inflammatory mediators, impacting tumor progression and response to therapies [51].
Specifically, SCFAs produced by Megasphaera and Lactobacillus, such as butyrate, acetate, and propionate, can act as histone deacetylase inhibitors, altering chromatin accessibility and modulating the expression of genes involved in proliferation and survival. The reduction of protective taxa like Actinomyces, which normally produce anti-inflammatory metabolites, may further exacerbate PI3K/AKT pathway activation, promoting a more proliferative and tumor-permissive microenvironment [52, 53].
CTNNB1
CTNNB1 mutations lead to the activation of the Wnt/β-catenin pathway which is crucial for cell proliferation and differentiation [54]. The gut microbiome plays a role in modulating this pathway. Specific taxa such as Fusobacterium and Akkermansia have been shown to impact Wnt signaling, immune infiltration, and tumor progression [56]. F. nucleatum can adhere to colonic epithelial cells via its FadA adhesin, activating β-catenin signaling and driving transcription of oncogenes that promote proliferation and survival. It also recruits myeloid cells and suppresses cytotoxic T-cell activity, creating a pro-tumorigenic immune microenvironment. On the other hand, Akkermansia muciniphila has been shown to promote proliferation and differentiation of intestinal stem cells by activating the Wnt/β-catenin pathway, while also maintaining epithelial integrity and moderating immune responses via the Treg/Th17 balance, supporting a healthy gut environment and potentially limiting tumor initiation and progression [55].
APC
Adenomatous Polyposis Coli (APC) mutations are among the most prevalent genetic alterations in EOCRC. Individuals carrying pathogenic APC mutations can develop hundreds to thousands of adenomatous polyps, leading to onset of CRC often before age 40 [56]. Pathogenic APC mutations, typically truncating variants, disrupt the β-catenin destruction complex, allowing nuclear accumulation of β-catenin and constitutive activation of the Wnt/β-catenin signaling that drives epithelial proliferation and survival [56, 57].
Microbial profiling of APC-mutated EOCRC demonstrated enrichment of Fusobacterium mortiferum and depletion of beneficial taxa such as Faecalibacterium prausnitzii and Bifidobacterium pseudocatenulatum [58]. F. mortiferum may exacerbate Wnt pathway activation through epithelial adherence and induction of local inflammatory cascades, reinforcing β-catenin-dependent proliferation. In contrast, F. prausnitzii and B. pseudocatenulatum support epithelial barrier integrity and immune homeostasis through SCFA production and suppression of proinflammatory signaling, including NF-κB-mediated pathways. Loss of these protective taxa in APC-mutated EOCRC may thus remove critical anti-inflammatory and immunoregulatory influences, shifting the mucosal environment toward sustained Wnt/β-catenin activation and tumor growth.
KRAS
KRAS mutations are found in 40% of CRC cases and promote persistent activation of oncogenic pathways such as MAPK/ERK and PI3K/AKT. This contributes to poor prognosis, tumor growth, and limited responsiveness to anti-EGFR therapies [59]. This persistent signaling promotes cellular proliferation, survival, and metabolic reprogramming, contributing to tumor progression, therapeutic resistance, and poor prognosis.
Microbiome profiling studies have revealed that KRAS-mutated tumors exhibit distinct compositional shifts within the gut microbiome, including enrichment of Fusobacterium, Clostridium, and Shewanella species [60]. Among these, F. nucleatum plays a central role in exacerbating KRAS-driven oncogenicity by activating TLR4-MYD88-NF-κB signaling to sustain pro-inflammatory cytokine and microRNA-21 expression, which supports tumor cell survival, invasion, and epithelial-mesenchymal transition.
Additionally, F. nucleatum also promotes immune evasion through Fap2-TIGIT-mediated inhibition of natural killer and cytotoxic T-cell activity [61]. Additional taxa, including Clostridium species and Shewanella, may further modulate tumor behavior through context-dependent effects on microbial metabolite production and oxidative stress [62, 63].
Clinical Therapies
Current therapies for EOCRC largely mirror that of LOCRC, including chemotherapy and targeted therapies for pMMR/MSS patients and immunotherapy for dMMR/MSI-H patients [4]. Importantly, survivorship factors are even more significant due to the impact of long-term side effects, including fertility issues, treatment-related toxicities, and unique tumor challenges which are not addressed by current treatments [70].
The gut microbiome is an emerging target for EOCRC clinical therapies and management. Distinct microbial patterns, such as the enrichment of Fusobacterium nucleatum, Bacteroides fragilis, and pks+ Escherichia coli, not only impact EOCRC pathogenesis, but influence therapy response. For example, Fusobacterium nucleatum has been seen to resist fluoropyrimidine-based chemotherapy, while other microbes, such as Akkermansia muciniphila has been linked to better responses with immune checkpoint inhibitors [74]. Such findings suggest that these microbial signatures can potentially be used as non-invasive biomarkers as predictors of immune response.
Early clinical studies find that prebiotics, probiotics, selective antibiotics, fecal microbiota transplantation (FMT), and dietary interventions show promise in tumor alteration, treatment efficacy enhancement, and treatment-related toxicity mitigation [75]. In the future, integration of microbiome research with tumor genomics and epigenetics can enable personalized EOCRC treatment. Ultimately, advancing these efforts may allow EOCRC clinical care to move from LOCRC borrowed strategies toward tailored approaches which improve both outcomes and survivorship for young patients.
Beyond treatment, the microbiome holds potential for public health and prevention. Because lifestyle and environmental choices play such a large role in EOCRC, early intervention can incorporate risk scoring. These screening guidelines can identify at-risk young individuals who may benefit from a colonoscopy or stool testing. Such approaches can shift the EOCRC approach from late detection to early prevention.
Future Directions
While we have improved our understanding of EOCRC risk factors, gaps remain in understanding how early-life exposures, antibiotic use, diet, and host-microbe interactions converge to shape the development of EOCRC [12]. In fact, the potential of the gut microbiome as a biomarker for early detection or as a therapeutic target, remains underexplored in this population. Studies on gut microbiome mediation of risks through inflammation remain cross-sectional, making it difficult to determine whether microbial changes precede tumorigenesis or arise because of disease. Longitudinal studies in at-risk young populations and EOCRC-focused studies into lifestyle and hereditary syndromes remain limited. Addressing these gaps is essential to clarify causal pathways and identify microbial targets for prevention and early detection.
Conclusion
EOCRC is an increasing worldwide health issue, presenting distinct clinical difficulties and unfavorable outcomes that highlight the necessity for greater insight beyond what is understood from late-onset disease. This review has underscored the importance of the gut microbiome in EOCRC, integrating current evidence on microbial profiles, metabolites, and bacterial genotoxins, while emphasizing how these elements relate to known risk factors like obesity, Lynch syndrome, inflammatory bowel disease, and lifestyle influences. Filling these gaps with longitudinal, age-stratified, and mechanistic studies will be essential for progressing from correlation to causations. In the end, promoting microbiome-centered research in EOCRC holds considerable potential to enhance early detection, inform prevention strategies, and support current therapies, with the main objective of alleviating the impact of this rising cancer in younger individuals.
References
- Morgan E, Arnold M, Gini A, Lorenzoni V, Cabasag CJ, et al. (2023) Global burden of colorectal cancer in 2020 and 2040: incidence and mortality estimate from GLOBOCAN. Gut. 72:338-344.
- Xiang Z, Luo P, Zhang X, Song Y, Zhang Z, et al. (2023) Oncotherapeutic strategies in early-onset colorectal cancer. Cancers (Basel).15:552.
- Dienstmann R, Salazar R, Tabernero J (2015) Personalizing Colon Cancer Adjuvant Therapy: Selecting Optimal Treatments for Individual Patients. J Clin Oncol. 33:1787-96.
- Xu K, Motiwala Z, Corona-Avila I, Makhanasa D, Alkahalifeh L, (2025) The gut microbiome and its multifaceted role in cancer metabolism, initiation, and progression: insights and therapeutic implications. Technol Cancer Res Treat. 24:15330338251331960.
- Gomes de Sousa R, Guerreiro CS, Santos I, Cravo M (2025) The knowledge gap in gut microbiome characterization in early-onset colorectal cancer patients: a systematic scoping review. Cancers (Basel).17:1863.
- Du M, Gu D, Xin J, Peters U, Song M, et al. (2023) Integrated multi-omics approach to distinct molecular characterization and classification of early-onset colorectal cancer. Cell Rep Med. 4:100974.
- Storandt MH, Rogen KR, Iyyangar A, Schnell RR, Mitchell JL, et al. (2025) Genomic landscapes of early-onset versus average-onset colorectal cancer populations. Cancers (Basel).17:836.
- Zhang J, Haines C, Watson AJM, Hart AR, Platt MJ, et al. (2019) Oral antibiotic use and risk of colorectal cancer in the United Kingdom, 1989–2012: a matched case– control study. Gut. 68:1971-1978.
- Zhang J, Haines C, Watson AJM, Hart AR, Platt MJ, et al. (2019) Oral antibiotic use and risk of colorectal cancer in the United Kingdom, 1989–2012: a matched case– control study. Gut. 68:1971-1978.
- Rebersek M. (2021) Gut microbiome and its role in colorectal cancer. BMC Cancer. 21:1325.
- Barot SV, Jandhyala DM, Chen R, Mena GE, Wang H, et al. (2024) Distinct intratumoral microbiome of young-onset and average-onset colorectal cancer. EBioMedicine. 100:105522.
- Xiong H, Huang X, Ye G, Zhang Y, Liu Y, et al. (2022) Gut microbiota displays alternative profiles in patients with early-onset vs late-onset colorectal cancer. Front Cell Infect Microbiol. 12:1036946.
- McDowell R, Hughes C, Hamnett A, Coleman HG, Cardwell CR, et al. (2022) Oral antibiotic use and early-onset colorectal cancer: Findings from a case-control study using a national clinical database. Br J Cancer. 127:766-776.
- Biragyn A, Luigi F (2018) Gut Dysbiosis: A Potential Link Between Increased Cancer Risk in Ageing and Inflammaging. Lancet Oncology. e295-e304.
- Rahman S, Becker SL, Patel RK, Martindale RG, Tsikitis VL, et al. (2023) Gut Microbial Metabolites and Its Impact on Human Health. Annals of Gastroenterology. 36: 360-368.
- Zhang Y, Chen R, Zhang D, Qi S, Liu Y, et al. (2023) Metabolite interactions between host and microbiota during health and disease: Which feeds the other? Br J Biomed Pharmacother. 160:114295.
- Gou H, Zeng R, Lau HCH, Yu J (2024) Gut microbial metabolites: Shaping future diagnosis and treatment against gastrointestinal cancer. Pharmacol Res. 208:107373.
- Pan H, Zhang L, Pang Y, Li J, Cheng Y, et al. (2025) Global burden, trends, and risk factors of early-onset and later-onset colorectal cancer: an analysis of the Global Burden of Disease Study 1990-2021, with projections to 2040: a population-based study. BMC Med. 25:486.
- Xu P, Tao Z, Yang H, Zhang C (2024) Obesity and early-onset colorectal cancer risk: emerging clinical evidence and biological mechanisms. Front Oncol. 14:1366544.
- Galasso L, Termite F, Mignini I, Esposto G, Borriello R, et al. (2025) Unraveling the Role of Fusobacterium nucleatum in Colorectal Cancer: Molecular Mechanisms and Pathogenic Insights. Cancers. 17:368.
- De Simone V, Franzè E, Ronchetti G, Colantoni A, Fantini MC, et al. (2015) Th17type cytokines, IL-6 and TNF-α synergistically activate STAT3 and NF-kB to promote colorectal cancer cell growth. Oncogene. 34:3493-503.
- Allen J, Rosendahl Huber A, Pleguezuelos-Manzano C, Puschhof J, Wu S, et al. (2022) Colon tumors in Enterotoxigenic Bacteroides fragilis (ETBF)-colonized mice do not display a unique mutational signature but instead possess host-dependent alterations in the APC gene. Microbiology Spectrum.10: e0105522.
- Pleguezuelos-Manzano C, Puschhof J, Rosendahl Huber A, Van Hoeck A, Wood HM, et al. (2020) Mutational signature in colorectal cancer caused by genotoxic pks+ E. coli. Nature. 580:269-273.
- Jiao Y, Ma Y (2025) Pesticides and Inflammatory Bowel Disease: A Critical Look at the Hidden Gaps in the Exposome-Disease Link. Gastroenterology.169:367.
- Wang X, Zheng Z, Yu D, Qiu X, Yang T, et al. (2025) Colorectal cancer in Lynch syndrome families: consequences of gene germline mutations and the gut microbiota. Orphanet J Rare Dis. 20:30.
- Lang M, Baumgartner M, Rożalska A, Frick A, Riva A, et al. (2020) Crypt residing bacteria and proximal colonic carcinogenesis in a mouse model of Lynch syndrome. Int J Cancer. 147:2316-2326.
- Qu S, Zheng Y, Huang Y, Feng Y, Xu K, et al. (2023) Excessive consumption of mucin by over-colonized Akkermansia muciniphila promotes intestinal barrier damage during malignant intestinal environment. Front Microbiol. 14:1111911.
- Sadeghi M, Mestivier D, Carbonnelle E, Benamouzig R, Khazaie K, et al. (2023) Loss of symbiotic and increase of virulent bacteria through microbial networks in Lynch syndrome colon carcinogenesis. Front Oncol. 13:1313735.
- Dejea CM, Fathi P, Craig JM, Boleij A, Taddese R, et al. (2018) Patients with familial adenomatous polyposis harbor colonic biofilms containing tumorigenic bacteria. Science. 359:592-597.
- Chen Y, Chen H, Hong J, Zou W, Fang JY, et al. (2017) Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell. 170:548-563.e16.
- Debesa-Tur G, Pérez-Brocal V, Ruiz-Ruiz S, Castillejo A, Latorre A, et al. (2021) Metagenomic analysis of formalin-fixed paraffin-embedded tumor and normal mucosa reveals differences in the microbiome of colorectal cancer patients. Sci Rep. 11:391.
- Ding T, Wu M, Zhao L, Liu H, Cao X, et al. (2025) Fusobacterium nucleatum downregulated MLH1 expression in colorectal cancer by activating the autophagy-lysosome pathway and is correlated with microsatellite instability and deficient mismatch repair. Front Oncol. 16:1586146.
- Yuan X, Lau H-C, Huang H, Hsueh C-Y, Gong H, et al. (2024) Integrative methylome and transcriptome analysis reveals epigenetic regulation of Fusobacterium nucleatum in laryngeal cancer. Microb Genom. 10:001221.
- Byrd DA, Fan W, Greathouse KL, Wu MC, Xie H, et al. (2023) The intratumor microbiome is associated with microsatellite instability. JNCI J Natl Cancer Inst. 115:989-993.
- Hamada T, Zhang X, Mima K, Bullman S, Sukawa Y, et al. (2018) Fusobacterium nucleatum in Colorectal Cancer Relates to Immune Response Differentially by Tumor Microsatellite Instability Status. Cancer Immunol Res. 6:1327-1336.
- Lucafò M, Curci D, Franzin M, Decorti G, Stocco G, et al. (2021) Inflammatory bowel disease and risk of colorectal cancer: An overview from pathophysiology to pharmacological prevention. Front Pharmacol. 12:772101.
- Zheng J, Sun Q, Zhang M, Liu C, Su Q, et al. (2024) Noninvasive, microbiome-based diagnosis of inflammatory bowel disease. Nat Med. 30:3555-67.
- Lucas C, Barnich N, Nguyen HTT (2017) Microbiota, inflammation and colorectal cancer. Int J Mol Sci. 18:1310.
- Cao Y, Oh J, Xue M, Huh WJ, Wang J, et al. (2022) Commensal microbiota from patients with inflammatory bowel disease produce genotoxic metabolites. Science. 378: eabm3233.
- Xia X, Wu WKK, Wong SH, Liu D, Kwong TNY, et al. (2020) Bacteria pathogens drive host colonic epithelial cell promoter hypermethylation of tumor suppressor genes in colorectal cancer. Microbiome. 8:108.
- Wang X, Allen TD, May RJ, Lightfoot S, Houchen CW, et al. (2008) Enterococcus faecalis induces aneuploidy and tetraploidy in colonic epithelial cells through a bystander effect. Cancer Res. 68:990-17.
- Lin R, Zhang Y, Chen L, Qi Y, He J, et al. (2020) The effects of cigarettes and alcohol on intestinal microbiota in healthy men. J Microbiol.58:926-937.
- Hur J, Otegbeye E, Joh HK, Nimptsch K, Ng K, et al. (2021) Sugar-sweetened beverage intake in adulthood and adolescence and risk of early-onset colorectal cancer among women. Gut. 70:2330-2336.
- Cao Y, Wu K, Mehta R, Drew DA, Song M, et al. (2016) Long-term use of antibiotics and risk of colorectal adenoma. Gut.gutjnl.67:672-678.
- Jiang F, Boakye D, Sun J, Wang L, Yu L, et al. (2023) Association between antibiotic use during early life and early‐onset colorectal cancer risk overall and according to polygenic risk and FUT2 genotypes. Int J Cancer.153:1602-1611.
- Li J, Pan Y, Guo F, Wang C, Liang L, et al. (2025) Patterns in genomic mutations among patients with early-onset colorectal cancer: an international, multicohort, observational study. Lancet Oncol. 26:1055-1066.
- Könönen E, Wade WG (2015) Actinomyces and related organisms in human infections. Clin Microbiol Rev.28:419-42.
- Bai D, Zhao J, Wang R, Du J, Zhou C, et al. (2024) Eubacterium coprostanoligenes alleviates chemotherapy-induced intestinal mucositis by enhancing intestinal mucus barrier. Acta Pharm Sin B.14:1677-1692.
- Chiang JYL, Ferrell JM (2020) Bile acid receptors FXR and TGR5 signaling in fatty liver diseases and therapy. Am J Physiol Gastrointest Liver Physiol.318(3): G554-G573.
- Wang H,Tang R, Jiang L,Jia Y. (2024) The role of PIK3CA gene mutations in colorectal cancer and the selection of treatment strategies. Front Pharmacol. 15:1494802.
- Yuan D, Tao Y, Wang H, Wang J, Cao Y, et al. (2022) A comprehensive analysis of the microbiota composition and host driver gene mutations in colorectal cancer. Invest New Drugs. 40:884-894.
- Markowiak-Kopeć P, Śliżewska K. (2020) The Effect of Probiotics on the Production of Short Chain Fatty Acids by Human Intestinal Microbiome. Nutrients. 12:1107.
- Verma A, Bhagchandani T, Rai A, Nikita, Sardarni UK, et al. (2024) Short-Chain Fatty Acid (SCFA) as a Connecting Link between Microbiota and Gut-Lung Axis-A Potential Therapeutic Intervention to Improve Lung Health. ACS Omega. 9:14648-14671.
- Marx O, Mankarious M, Yochum G (2023) Molecular genetics of early-onset colorectal cancer. World J Biol Chem.14:13-27.
- Ma X, Li M, Zhang Y, Xu T, Zhou X, et al. (2025) Akkermansia muciniphila identified as key strain to alleviate gut barrier injury through Wnt signaling pathway. eLife.12:RP92906.
- Hankey W, Frankel WL, Groden J (2018) Functions of the APC tumor suppressor protein dependent and independent of canonical WNT signaling: implications for therapeutic targeting. Cancer Metastasis Rev. 37:159-172.
- Zhang D, Ni QQ, Wang SY, He WF, Hong ZX, et al. (2024) APC mutations disrupt βcatenin destruction complex condensates organized by Axin phase separation. Cell Mol Life Sci.81:57.
- Liang S, Mao Y, Liao M, Xu Y, Chen Y, et al. (2020) Gut microbiome associated with APC gene mutation in patients with intestinal adenomatous polyps. Int J Biol Sci.16:135-146.
- Takeda M, Yoshida S, Inoue T, Sekido Y, Hata T, et al. (2025) The Role of KRAS Mutations in Colorectal Cancer: Biological Insights, Clinical Implications, and Future Therapeutic Perspectives. Cancers. 17:428.
- Huang Z, Huang X, Huang Y, Liang K, Chen L, et al. (2024) Identification of KRAS mutation-associated gut microbiota in colorectal cancer and construction of predictive machine learning model. Navarathna DH, editor. Microbiol Spectr. 12: e0272023.
- Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, et al. (2013) Fusobacterium nucleatum Promotes Colorectal Carcinogenesis by Modulating E-Cadherin/β-Catenin Signaling via its FadA Adhesin. Cell Host Microbe.14:195-206.
- Wang Y, Huang D, Wang R, Yin J, Pei J, et al. (2025) Microbial metabolites in tumor metabolic reprogramming and immunotherapy: new insights. Front. Cell. Infect. Microbiol.15:1706040.
- Yang L, Wu H, Zhao Y, Tan X, Wei Y, et al. (2024) Shewanella oneidensis MR-1 dissimilatory reduction of ferrihydrite to highly enhance mineral transformation and reactive oxygen species production in redox-fluctuating environments. Chemosphere.352:141364.
- Sunami E, de Maat M, Vu A, Turner RR, Hoon DSB, et al. (2011) LINE-1 hypomethylation during primary colon cancer progression. PloS One.6:e18884.
- Antelo M, Balaguer F, Shia J, Shen Y, Hur K, et al. (2012) A High Degree of LINE-1 Hypomethylation Is a Unique Feature of Early-Onset Colorectal Cancer. PLoS ONE.7: e45357.
- Xiong H, Wang J, Chang Z, Hu H, Yuan Z, et al. (2022) Gut microbiota display alternative profiles in patients with early-onset colorectal cancer. Front Cell Infect Microbiol.12:1036946.
- D’Aquila P, Carelli LL, De Rango F, Passarino G, Bellizzi D, et al. (2020) Gut microbiota as important mediator between diet and DNA methylation and histone modifications in the host. Nutrients. 12:597.
- Li Y, Chen W, Yu J, Zhang S, Wang J, et al. (2022) Multi-omics analyses reveal multimodal Fusobacterium DNA damage perturbations in colorectal cancer. Cancer Res. 82: 6212.
- Yang Y, Weng W, Peng J, Hong L, Yang L, et al. (2017) Fusobacterium nucleatum increases proliferation of colorectal cancer cells by activating TLR4 signalling to NF-κB, upregulating miR-21, and downregulating RASA1. Gastroenterology.152:851-66.e24.
- Grothey A, Sobrero AF, Shields AF, Yoshino T, Paul J, et al. (2018) Duration of Adjuvant Chemotherapy for Stage III Colon Cancer. N Engl J Med. 378:1177-1188.
- Zhang X, Wu T, Cai X, Dong J, Xia C, et al. (2022) Neoadjuvant Immunotherapy for MSIH/dMMR Locally Advanced Colorectal Cancer: New Strategies and Unveiled Opportunities. Front Immunol. 13:795972.
- André T, Shiu KK, Kim TW, Jensen BV, Jensen LH, et al. (2020) Pembrolizumab in Microsatellite-Instability–High Advanced Colorectal Cancer. N Engl J Med.383:2207-2218.
- Kumar A, Gautam V, Sandhu A, Rawat K, Sharma A, et al. (2023) Current and emerging therapeutic approaches for colorectal cancer: A comprehensive review. World J Gastrointest Surg.15:495-519.
- Eslami M, Naderian R, Bahar A, Babaeizad A, Rezanavaz Gheshlagh S, et al. (2025) Microbiota as diagnostic biomarkers: advancing early cancer detection and personalized therapeutic approaches through microbiome profiling. Front Immunol.16:1559480.
- Lei W, Zhou K, Lei Y, Li Q, Zhu H, et al. (2025) Gut microbiota shapes cancer immunotherapy responses. Npj Biofilms Microbiomes.11:143.
- Gomes De Sousa R, Guerreiro CS, Santos I, Cravo M (2025) The Knowledge Gap in Gut Microbiome Characterization in Early-Onset Colorectal Cancer Patients: A Systematic Scoping Review. Cancers. 17:1863.
© by the Authors & Gavin Publishers. This is an Open Access Journal Article Published Under Attribution-Share Alike CC BY-SA: Creative Commons Attribution-Share Alike 4.0 International License. Read More About Open Access Policy.