Biomarker Proliferation, Biological Overlap, and the Specificity Illusion: An Editorial on Interpretation, Validation, and Use
by George B. Stefano*
Mind-Cell LLC, 841 E. Fort Ave, B-411, Baltimore, MD 21230, USA
*Corresponding Author: George B. Stefano, Mind-Cell LLC, 841 E. Fort Ave, B-411, Baltimore, MD 21230, USA
Received Date: 20 February 2026
Accepted Date: 23 February 2026
Published Date: 25 February 2026
Citation: George B. Stefano (2026) Biomarker Proliferation, Biological Overlap, and the Specificity Illusion: An Editorial on Interpretation, Validation, and Use. J Surg 11: 11575 DOI: https://doi.org/10.29011/2575-9760.011575
Abstract
The biomedical literature has seen a tremendous growth in the field of biomarker research, driven by improvements in high-throughput measurement, computational analytics, and translational research. While this has helped to provide important mechanistic insights and novel clinical applications, it has also seen an explosion of biomarker research that often blurs the distinction between statistical association and disease specificity. Many of the well-characterized biomarkers represent processes that are commonly involved across various disease states, including those that are inflammatory, metabolically stressful, immunologically activated, or injury-related. As a consequence, outcome-based research may inadvertently overlook the presence of the same biomarker across related but distinct disease states. This editorial attempts to identify the biological and methodological underpinnings for the nonspecificity of biomarkers, the advantages and disadvantages associated with the current discovery approach, and suggests a context-of-use-driven approach to validation-based research to improve the overall utility and reproducibility of biomarkers.
Keywords: Biomarkers; Inflammation; Mitochondrial Dysfunction; Post-Acute Covid-19 Syndrome; Systems Biology; Translational Medical Research
Editorial Discussion
The fast development of biomarker research is an indication of significant scientific progress. New high-dimensional -omics technologies, immunoassays, and machine learning approaches have enabled biological signals to be quantified and associated with disease risk, progression, or treatment response. New and updated regulatory and translational approaches now formally acknowledge different types of biomarkers, including diagnostic, prognostic, predictive, pharmacodynamic, and monitoring biomarkers, each of which has different criteria for evidence [1]. Simultaneously, significant advances in blood and fluid biomarkers for neurodegenerative diseases represent the potential and complexity of current biomarker research, particularly when moving from research to the clinic [2,3]. In this environment, biomarkers have been incredibly powerful tools for dissecting heterogeneous populations, finding converging biological mechanisms, and accelerating therapeutic development. At the same time, this growth has highlighted an underlying limitation of biomarker interpretation: many biomarkers may be biologically valid but nonspecific. One of the most common trends in the literature is the finding of a biomarker that is very tightly associated with disease but primarily reflects a general physiological condition. Inflammatory biomarkers, such as interleukin-6 (IL-6) and C-reactive protein, reliably measure the presence of systemic immune activation and stress response but are elevated across a wide range of conditions, including infection, autoimmune disease, cardiovascular disease, neuropsychiatric disorders, and psychosocial stress [4]. They may identify the presence of a shared biological response, but they do not define disease mechanisms. This problem is exacerbated in complex disorders with multiple contributing factors, in which symptom overlap and etiologic heterogeneity are common. In psychiatric disorders, neurodegeneration, metabolic disease, and oncology, biomarkers often reflect common downstream effects such as mitochondrial dysfunction, oxidative stress, neuroendocrine–immune coupling, and tissue remodeling rather than disorder-specific etiologies. Even with the use of sophisticated approaches such as polygenic risk scores, which are informative at the population level, substantial challenges remain with clinical specificity, interpretation, and translation to the individual patient [5]. Indeed, even when robust methods demonstrate differences between a disease group and healthy controls, the presence and magnitude of similar effects in related clinical populations may remain unknown. This can inadvertently lead to a “specificity illusion,” in which overall biological relevance is conflated with diagnostic specificity [6]. Methodologic factors also contribute to this issue. Biomarker discovery studies frequently face limitations related to small sample sizes, high dimensionality, and flexible analytical pipelines, increasing the risk of false discovery and overfitting rather than identification of true biological effects [7,8]. Meta-research has consistently demonstrated that these vulnerabilities are systemic, particularly in early-phase translational research [9,10]. In recognition of these challenges, reporting and study-design guidelines have been established. STARD for diagnostic accuracy studies, TRIPOD for prediction models, and REMARK for prognostic tumor markers emphasize transparent reporting of cohort selection, assay handling, statistical modeling, and validation strategies [11-13]. Design approaches such as prospective sample collection and blinded evaluation aim to minimize bias and improve generalizability [14]. However, even in established fields such as neurodegenerative biomarker research, translation to clinical utility remains inconsistent, underscoring the need for rigorous adherence to these standards [3]. Importantly, biomarker nonspecificity should not be equated with failure. In appropriate contexts, nonspecific biomarkers can illuminate shared biology across disease states rather than obscure it. Increasingly, it is recognized that biomarker panels, rather than single markers, are more likely to identify disease-relevant biological patterns while accounting for nonspecificity [2]. The critical issue is therefore not novelty or apparent specificity, but clarity regarding whether a biomarker serves mechanistic, stratification, monitoring, or discriminative functions (Figure 1).
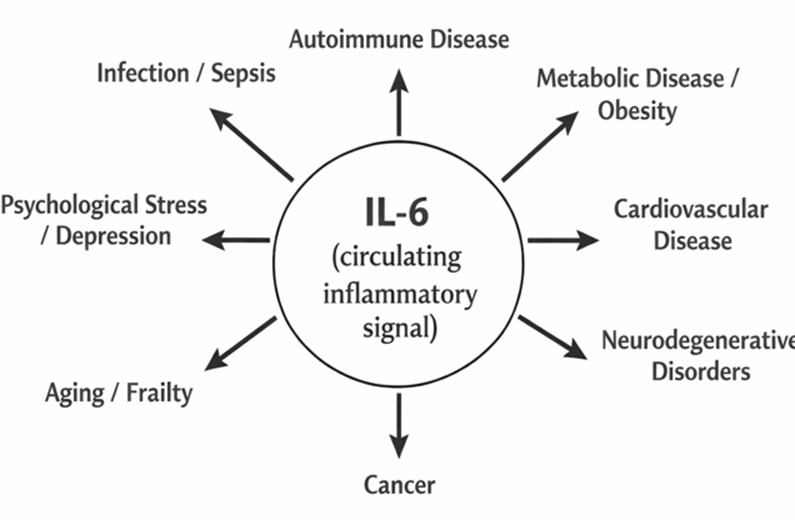
Figure 1: Interleukin-6 (IL-6) as a non-specific biomarker. The figure portrays IL-6 as the key circulating signal for inflammation that connects to a variety of disparate diseases, ranging from infectious diseases to autoimmune diseases, metabolic diseases, cardiovascular diseases, neurodegenerative diseases, cancer, psychological stress, and aging. The balanced presentation of all these diseases indicates that IL-6, as a biomarker for inflammation, represents a generalized systemic response to inflammation or stress and, as such, does not point to any particular disease process or pathology. Thus, IL-6, by itself, represents a non-specific biomarker for diseases in general.
Illustrative Example: Mitochondrial Damage Markers in Long COVID
A recent Molecular Psychiatry study from the EPILOC Phase 2 cohort provides a clear illustration of these principles. In a nested case–control analysis of 228 adults (128 with post-acute sequelae of COVID-19 and 100 recovered controls), Matits et al. quantified relative Circulating Cell-Free Mitochondrial DNA (ccf-mtDNA), total cell-free DNA, and inflammatory markers, relating these measures to cognitive performance, verbal memory, and psychological distress using principal component analysis [15]. Individuals with long COVID demonstrated reduced cognitive performance, increased psychological distress, elevated C-reactive protein, and lower relative ccf-mtDNA compared with recovered controls. Across the full cohort, relative ccf-mtDNA correlated positively with general cognition and inversely with inflammatory burden. Mediation analyses suggested that altered ccf-mtDNA may partially link inflammatory differences between groups, although these effects were attenuated after adjustment for demographic and clinical confounders [15]. This pattern closely mirrors long-standing observations with IL-6: elevations are biologically meaningful yet insufficiently specific to define a discrete disease process when considered in isolation. Accordingly, the EPILOC findings do not establish ccf-mtDNA as a disease-specific biomarker for long COVID, but rather place mitochondrial damage markers within the same interpretive category as IL-6—highly sensitive indicators of shared biological stress.
Alignment With the IL-6 Nonspecific Biomarker Paradigm
IL-6 has been associated with acute COVID-19 severity, persistent post-COVID symptoms, depression, cognitive impairment, metabolic disease, and neurodegeneration. Its primary interpretive value lies in signaling systemic immune activation rather than distinguishing underlying etiologies. The EPILOC study reinforces this principle by demonstrating that inflammation-related signals gain explanatory value only when embedded within a broader biological framework. Within this framework, mitochondrial damage markers operate both upstream and downstream of cytokine signaling. Mitochondrial dysfunction promotes inflammatory activation, while sustained inflammation further impairs mitochondrial quality-control processes. Circulating mitochondrial DNA is particularly informative because it lies at the intersection of metabolism and immunity, functioning both as a bioenergetic indicator and as a damage-associated molecular pattern [16,17]. Thus, ccf-mtDNA should be interpreted not as a disease signature but as a contextual biomarker that situates inflammatory signals, such as IL-6, within a coherent systems-biology model.
Mitochondria as Targets and Mediators in COVID-19
Independent evidence supports this interpretation. SARS-CoV-2 has been shown to disrupt mitochondrial signaling, redox balance, and mitochondrial DNA dynamics [18,19]. Integrative models linking microglial activation, impaired mitophagy, and altered mitochondrial DNA provide mechanistic continuity between viral infection and long-term neuropsychiatric manifestations [20]. In this context, reduced relative ccf-mtDNA in long COVID may reflect maladaptive mitochondrial quality-control mechanisms rather than simple cellular injury, consistent with observed neuropsychiatric outcomes.
Pros and Cons of the Current Biomarker Landscape
The expanding biomarker repertoire has advanced mechanistic understanding, interdisciplinary integration, and the efficiency of clinical research. Biomarkers enable patient stratification, longitudinal monitoring, and hypothesis testing across biological scales, and when applied within a clearly defined context of use, they can meaningfully inform both experimental and clinical decision-making [1]. Conversely, limitations related to marker specificity, study population selection, and inadequate comparison with competing diagnoses continue to constrain reproducibility and clinical translation. Failure to benchmark biomarker performance against relevant disease comparators, rather than healthy controls alone, remains a major barrier to implementation [8,21,5].
Conclusion
The exponential growth of biomarker research reflects both technological success and conceptual risk. Concerns that many focused outcome studies fail to recognize shared biomarker expression across related disorders are well supported by biological principles and empirical evidence. The EPILOC Phase 2 study illustrates how mitochondrial damage markers—like IL-6—can clarify shared biological stress without implying diagnostic specificity. Future progress will depend less on discovering new biomarkers and more on rigorously defining context of use, testing signals against appropriate comparators, and embedding biomarkers within interconnected biological systems. Only through this shift can biomarkers fulfill their promise as durable instruments of biomedical insight rather than transient statistical artifacts.
Author Contributions: Single author
Ethics Approval and Consent to Participate: Not applicable.
Acknowledgment: ChatGPT 5.2 was used for information organization and figure generation.
Funding: This research received no external funding.
Conflict of Interest: The author declares no conflict of interest.
References
- Cagney DN, Sul J, Huang RY (2018) The FDA–NIH Biomarkers, EndpointS, and other Tools (BEST) resource. Neuro-Oncology 20 1162-1172.
- Hampel H, Hu Y, Cummings J (2023) Blood-based biomarkers for Alzheimer’s disease: Current state and future use. Neuron 111: 2781-2799.
- Zetterberg H, Blennow K (2024) Fluid biomarkers for neurodegenerative disease. Nat. Rev. Neurol 20: 25-45.
- Del Giudice M, Gangestad SW (2018) Rethinking IL-6 and CRP. Brain Behav. Immun 70: 61-75.
- Martínez-Minguet D, Simón AG (2025) Challenges in clinical translation of polygenic risk scores. Genet. Med 27: 100309.
- Abi-Dargham A, Horga G, Miller EJ (2023) Candidate biomarkers in psychiatric disorders. World Psychiatry 22: 199-231.
- Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate. J. R. Stat. Soc. B 57: 289-300.
- Ioannidis JPA (2005) Why most published research findings are false. PLoS Med 2: e124.
- Ransohoff DF (2004) Rules of evidence for cancer molecular-marker discovery. Nat. Rev. Cancer 4 309-314.
- Ransohoff DF (2005) Bias as a threat to validity in molecular-marker research. Nat. Rev. Cancer 5: 142-149.
- Altman DG, McShane LM, Sauerbrei W, Taube SE (2012) REMARK guidelines. PLoS Med 9: e1001216.
- Bossuyt PM, Reitsma JB, Bruns DE (2015) STARD 2015. BMJ 351: h5527.
- Collins GS, Reitsma JB, Altman DG, Moons KGM (2015) TRIPOD statement. Ann. Intern. Med. 162: 55-63.
- Pepe MS, Feng Z, Janes H, Bossuyt PM, Potter JD (2008) Pivotal evaluation of a biomarker. J. Natl. Cancer Inst 100: 1432-1438.
- Matits L, Schellenberg J, Mack M (2026) Circulating mitochondrial and cellular damage markers in long COVID. Mol. Psychiatry.
- Esch T, Stefano GB, Ptacek R, Kream RM (2020) Blood-borne mitochondria and inflammation. Med. Sci. Monit 26: e924337.
- Stefano GB, Kream RM (2021) Mitochondrial DNA heteroplasmy and COVID-19 neurological disorders. Cell. Mol. Neurobiol 41: 1-10.
- Wang F, Kream RM, Stefano GB (2020) Long-term sequelae of COVID-19. Med. Sci. Monit 26: e928996.
- Stefano GB, Büttiker P, Weissenberger S (2021) Pathogenesis of long-term neuropsychiatric COVID-19. Med. Sci. Monit27: e933015.
- Stefano GB, Büttiker P, Weissenberger S (2023) Independent and sensory human mitochondrial functions. Front. Cell. Infect. Microbiol 13: 1130197.
- Kern SE (2012) Why your new cancer biomarker may never work. Cancer Res 72: 6097-6101.
© by the Authors & Gavin Publishers. This is an Open Access Journal Article Published Under Attribution-Share Alike CC BY-SA: Creative Commons Attribution-Share Alike 4.0 International License. Read More About Open Access Policy.