An Unexpected Finding in Cell Free DNA ddPCR Testing for BRAF V600E
by John A. Puskas, Aaron D. Bossler, Mark D. Foster, Yanet Prieto Carratala, Dahui Qin*
Department of Pathology/Molecular Diagnostic Laboratory/Moffitt Cancer Center, Tampa, Florida, 33647, USA
*Corresponding author: Dahui Qin, Department of Pathology/Molecular Diagnostic Laboratory/Moffitt Cancer Center, Tampa, Florida, 33647, USA
Received Date: 09 April 2026
Accepted Date: 15 April 2026
Published Date: 17 April 2026
Citation: Puskas JA, Bossler AD, Foster MD, Carratala YP, Qin D. (2026). An Unexpected Finding in Cell Free DNA ddPCR Testing for BRAF V600E. Ann Case Report. 11: 2595. DOI: https://doi.org/10.29011/2574-7754.102595
Abstract
A quantitative detection of BRAF V600E mutation in cell free DNA of plasma using ddPCR has been used in testing liquid biopsy and in follow-up tests of melanoma patients to monitor melanoma recurrence and to quantitatively measure minimal residual disease. In our CLIA certified laboratory, 364 samples have been tested in a 5-year time span. Eighty-five BRAF V600E were detected with various allele frequences, from 0.23% to 65.40%. The raw data of ddPCR is usually depicted in a 2D plot with 4 quadrants, similar to flowcytometry. BRAF codon 600 is interrogated in a droplet PCR reaction and generates a read from each droplet. The read is represented by a dot on the 2D plot. The positive dots are distributed in the upper left and right quadrants. The negative dots, the lower left and right quadrants. Among the 85 positive cases, we have encountered some unexpected findings from four cases. On the 2D plots, the positive dots of these cases are unexpectedly distributed on the border of the upper left and lower left quadrants, an atypical 2D plot pattern. Since some of the signal dots are above the cut off, the test results are output as positive. The four cases were re-tested using pyrosequencing and/or next generation sequencing. The results turn out to be BRAF V600K instead of BRAF V600E. Our ddPCR assay is designed to detect BRAF V600E mutation, but not for BRAF V600K. Such an unexpected finding prompts us to review the raw data on the 2D plot routinely. If an atypical 2D plot pattern is seen, we defer to an orthogonal assay instead of issuing a report of BRAF V600E. If the BRAF V600K mutation is verified by an orthogonal assay, a BRAF V600K positive report can be issued, citing the orthogonal test result.
Keywords: BRAF V600E; BRAF V600K; Digital PCR
Introduction
Liquid biopsy has been widely used in molecular diagnostic tests. Liquid biopsy is usually defined as obtaining the plasma and extracting circulating cell free tumor DNA (ctDNA) from the plasma for molecular tests [1]. This concept is expanding into the ctDNA from other body fluids [2, 3]. The ctDNA BRAF V600E test has been validated and used in patients with different diagnosis [4-6]. The BRAF V600E mutation is the most common driver mutation in Melanoma [7, 8]. The BRAF V600K mutation is the second most common driver mutation among BRAF mutated melanoma cases, followed by BRAF V600R, V600D etc. [8]. The ctDNA has been widely used to detect BRAF mutations in melanoma patients [9-13]. It has been specifically useful in follow-up tests to monitor patients’ response to treatment and to monitor the minimal residual disease and potential recurrence [14-18]. There are different methods in detecting BRAF mutations in ctDNA. Droplet Digital PCR (ddPCR) is one of the common methods [19-23]. The ddPCR assay has been used in our laboratory to detect BRAF V600E from ctDNA. In our practice, apart from detecting BRAF V600E mutation, this method occasionally generates an unexpected data, which is different from the data of a BRAF V600E negative sample and it is also different from the data of BRAF V600E positive samples. The investigation of such cases is reported here.
Case Report
Materials and Methods: Plasma ctDNA was extracted using the QIAmp Circulating Nucleic Acid kit (Qiagen, Inc. cat. #55114) according to the suggested protocol. For each sample, 30ng of ctDNA was used in ddPCR testing. The ddPCR was performed using the BRAF V600E ddPCR (FAM + HEX) probe assay (BioRad Inc.) following the manufacturer’s protocol. The BioRad QX200 Automated Droplet Generator and Reader, C1000 thermocycler, and QuantaSoft analysis software were used in the testing process. The ddPCR quantitates BRAF V600E mutant and wild type allele copy number and depicts the data in a two-dimensional (2D) dot plot similar to that of flow cytometry data. The mutant channel threshold (MCT) is set with water and BRAF wild type DNA negative controls (Figure 1). Signal dots above MCT are considered positive and results are reported as copies/ml plasma with the variant allele frequency.
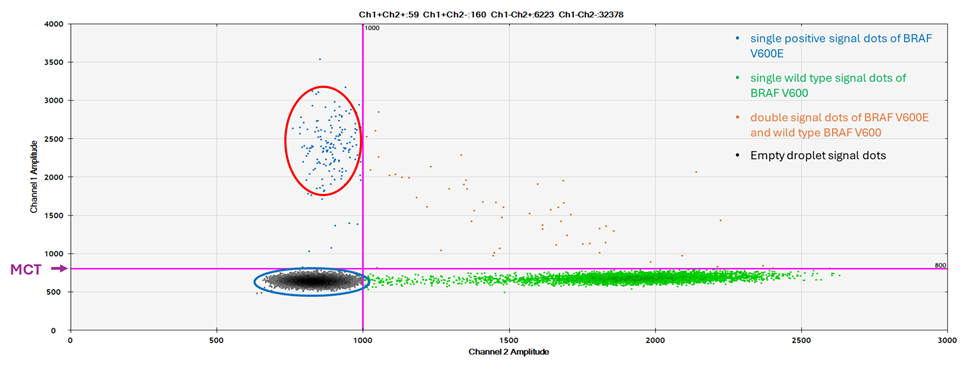
Figure 1: A 2D plot of a BRAF V600E positive result of ddPCR assay. The red circle is the area where the typical single positive signal dots of BRAF V600E are located. The blue circle is the area where the typical single signal dots of wild type BRAF are located. The orange dots in the upper right quadrant are droplets containing both BRAF V600E and BRAF wild type signals.
Results
Normally, the result of a tested sample is either positive for BRAF V600E or BRAF V600 wild type. In the normal BRAF V600 E positive cases, the typical single positive signals are the dots located in the left upper quadrant of the 2D plot above the MCT (Figure 1, red circle). The typical single signals of wild type BRAF are the dots located in the right lower quadrant of the 2D plot. The negative signals from empty droplets (neither BRAF V600E nor BRAF V600 wild type) are the dots located in the left lower quadrant (Figure 1, blue circle).
However, among 364 tests, the data from 4 cases showed an unexpected dot plot pattern, which has a group of signal dots in between the upper left quadrant and the lower left quadrant of the 2D dot plot. These signal dots are a distinct group of dots, which are located around the MCT, the signal intensity is significantly lower (seen in Figure 2, red circle) than that of normal BRAF V600E positive signal dots (seen in Figure1, red circle). However, these signal dots are separated from the negative signal dots from empty droplets (Figure 2, blue circle). Since some of the signal dots are above the cut off, the test results are output as positive for V600E.
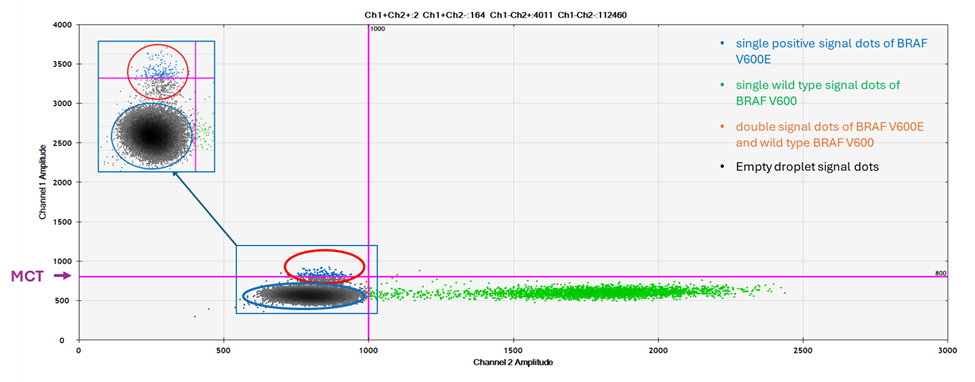
Figure 2: A 2D plot of unexpected finding. The red circle is the unexpected finding. The signal intensity is higher than the negative signals, but significantly lower than the normal BRAF V600E positive signal seen in Figure 1. The insert in the upper left quadrant is amplified along the Y axial to show that the red circle includes a distinct signal dot population, which is separated from that in blue circle.
A patients’ chart review showed that one case had a subcutaneous nodule of malignant melanoma tested positive for BRAF V600K. One case was negative for BRAF V600E on a BRAF V600E specific immunohistochemistry (IHC) test. Another two cases had no prior BRAF testing. We also had a BRAF V600R case, which had similar 2D plot pattern as seen in (Figure 2).
The ctDNA of all 4 samples were tested using pyrosequencing and/or NGS. All 4 samples are positive for the BRAF V600K mutation (Figure 3, panel A and B).
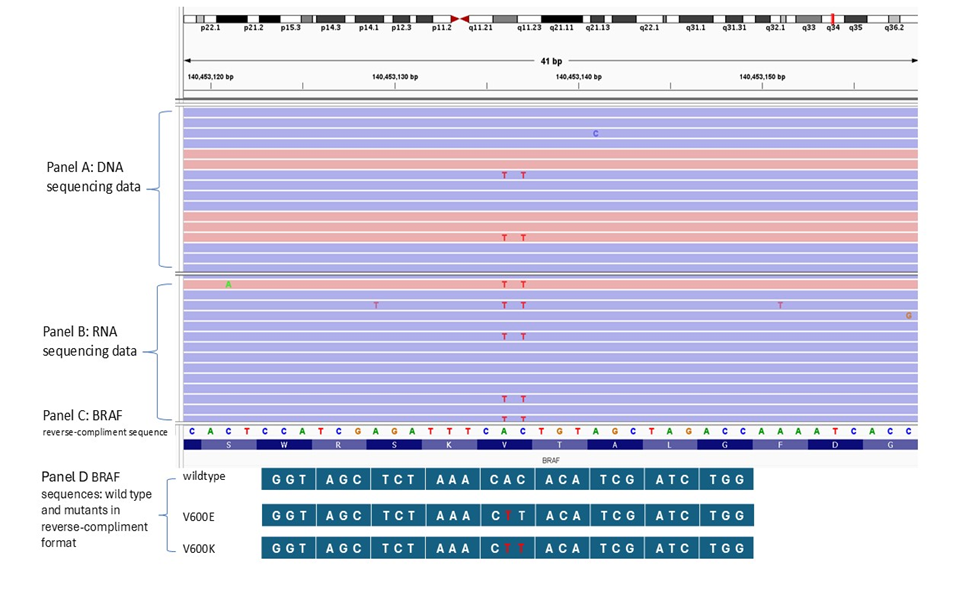
Figure 3: NGS result of a V600K sample and the sequences of BRAF wild type and mutants. Panel A is NGS sequencing result from the DNA of a V600K. Panel B is NGS sequencing result from the RNA of the V600K. Panel C is the BRAF reference sequencing used in NGS in a reverse-compliment format. Panel D is BRAF sequences of wild type and mutants in a reverse-compliment format.
Discussion
BRAF mutations are important driver mutations in melanoma [8]. The ddPCR assay for BRAF V600E is a highly sensitive assay in detecting ctDNA BRAF V600E. The lower detection limit of our assay is 0.23%. It also has very good specificity. This assay was designed to test BRAF V600E. Therefore, the samples used to evaluate the lower detection limit, the sensitivity and specificity are the samples either BRAF V600E positive or negative samples. However, in our clinical practice, some patients’ BRAF V600 mutation history is not certain. This leads to a possibility of some BRAF V600K and The unexpected findings present an unusual signal pattern on the 2D plot. Since some of the signal dots are above the cut off and the assay is designed for BRAF V600E detection, the test results are output as positive for V600E. The orthogonal tests show that these samples are actually positive for BRAF V600K. Portions of this work were presented in both abstract and poster form at the 2022 annual meeting of the Association for Molecular Pathology, November 3-5, 2022, Phoenix, AZ [24]. We believe that the unexpected findings are due to following causes. Since the assay is designed to detect the specific mutation of BRAF V600E, it is conceivable the assay has used the oligo specific PCR primers. The wild type sequence of BRAF around codon 600 in reverse-compliment format is GGTAGCTCTAAACACACATCGATCTGG, with the codon 600 bolded (also seen in Figure 3, panel D). The sequence of BRAF V600E is GGTAGCTCTAAACTCACATCGATCTGG with the mutated nucleotide in red (also seen in Figure 3, panel D). The primer specificity between the wild type and BRAF V600E will have to depend on the nucleotide difference at position of c.1799 of codon 600, where the wild type is an ‘A’ in a reverse-compliment format and BRAF V600E is a ‘T’. By convention, this should usually be the last nucleotide for primer design. While this works well for the specificity between the wild type and BRAF V600E mutant, it may pose a challenge for the primer specificity between BRAF V600E and BRAF V600K. The sequence around the BRAF V600K is GGTAGCTCTAAACTCTTATCGATCTGG with the mutated nucleotides in red (also seen in Figure 3, panel D). At the position c.1799, both BRAF V600E and BRAF V600K have the same nucleotide. Since it should be the last nucleotide of the BRAF V600E primer, it raises the possibility of its non-specific binding to BRAF V600K sequence though maybe at a significantly lower level. Since the ddPCR is a highly sensitive assay, such a low level of non-specific primer binding may generate a low level of ‘positive’ signal. This could explain why our BARF V600E specific assay generates a low positive signal in BRAF V600K samples. A similar phenomenon has been reported in ddPCR assay for EGFR, where one mismatched nucleotide could significantly change the signal pattern [25].
This unexpected finding reminds us that not all signals above the cut off are BRAF V600E. It is important to review the 2D plot for signal dot pattern. If the pattern is different from that of typical BRAF V600E, an orthogonal method needs to be used to verify if the case will be BRAF V600E or other BRAF V600 mutations. The good news is that Bio-Rad has developed assays for BRAF V600K, V600R and BRAF wild type, which are certainly helpful in testing and verifying non V600E samples.
Author contributions
JP: Data analysis, Writing-original draft, Writing-review & editing. AB: data analysis and manuscript-review & editing. MF: Data generating, Data analysis and manuscript-review & editing. YC: Data generating, Data analysis and manuscript-review & editing. DQ: Conceptualization, data analysis, Writing-original draft and Writing-review & editing.
Conflicts of interest
The authors declare that they have no conflicts of interest.
Ethical approval
Not required. This manuscript is focusing on de-identified sample testing technology and technical data analysis. The ethical approval is Not required per our institutional standards, which authorizes the waiver for this de-identified sample study.
Consent to participate
This manuscript is focusing on de-identified sample testing technology and technical data analysis. The consent is Not required per our institutional standards, which authorizes the waiver for this de-identified sample study.
Consent to publication
Not applicable.
Availability of data and materials
Not applicable.
Funding
Not applicable.
References
- Alidousty C, Brandes D, Heydt C, Wagener S, Wittersheim M, Schäfer SC, et al. (2017). Comparison of Blood Collection Tubes from Three Different Manufacturers for the Collection of Cell-Free DNA for Liquid Biopsy Mutation Testing. J Mol Diagn. 19: p. 801-804.
- Greuter L, Frank N, Guzman R, Soleman J. (2022). The Clinical Applications of Liquid Biopsies in Pediatric Brain Tumors: A Systematic Literature Review. Cancers (Basel). 14: 2683.
- Nikanjam M, Kato S, Kurzrock R. (2022). Liquid biopsy: current technology and clinical applications. J Hematol Oncol. 15: p. 131.
- Moradi-Marjaneh R, Asgharzadeh F, Khordad E, Marjaneh MM. (2021). The Clinical Impact of Quantitative Cell-free DNA, KRAS, and BRAF Mutations on Response to Anti-EGFR Treatment in Patients with Metastatic Colorectal Cancer. Curr Pharm Des. 27: p. 942-952.
- van Helden EJ, Anjus L, van der Houven van Oordt CWM, Heideman DAM, Boon E, et al. (2019). RAS and BRAF mutations in cell-free DNA are predictive for outcome of cetuximab monotherapy in patients with tissue-tested RAS wild-type advanced colorectal cancer. Mol Oncol, 13: p. 2361-2374.
- Arnolda R, Howlett K, Chan T, Raleigh J, Hatzimihalis A, et al. (2022). Clinical validation and implementation of droplet digital PCR for the detection of BRAF mutations from cell-free DNA. Pathology, 54: p. 772-778.
- Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, et al. (2011). Improved survival with vemurafenib in melanoma with BRAF V600E mutation. The New England journal of medicine. 364: p. 2507-2516.
- Castellani G, Buccarelli M, Arasi MB, Rossi S, Pisanu ME, et al. (2023). BRAF Mutations in Melanoma: Biological Aspects, Therapeutic Implications, and Circulating Biomarkers. Cancers (Basel), 15: 4026.
- Bos MK, Kraan J, Starmans MPA, Helmijr JCA, Verschoor N, et al. (2024). Comprehensive characterization of circulating tumor cells and cell-free DNA in patients with metastatic melanoma. Mol Oncol, 18: 2770-2782.
- Varaljai R, S Elouali, SS Lueong, K Wistuba-Hamprecht, Seremet T, et al. (2021). The predictive and prognostic significance of cell-free DNA concentration in melanoma. J Eur Acad Dermatol Venereol. 35: p. 387-395.
- Csoma SL, Madarász K, Yi CCC, Emri G, Bedekovics J, et al. (2023). Correlation Analyses between Histological Staging and Molecular Alterations in Tumor-Derived and Cell-Free DNA of Early-Stage Primary Cutaneous Melanoma. Cancers (Basel). 15: 5141.
- Rutkowski P, Pauwels P, Kerger J, Jacobs B, Maertens G, et al. (2021). Characterization and Clinical Utility of BRAF(V600) Mutation Detection Using Cell-Free DNA in Patients with Advanced Melanoma. Cancers (Basel), 13: 3591.
- Pinzani P, Salvianti F, Zaccara S, Massi D, De Giorgi V, et al. (2011). Circulating cell-free DNA in plasma of melanoma patients: qualitative and quantitative considerations. Clin Chim Acta, 412: 2141-2145.
- Gangadhar TC, Savitch SL, Yee SS, Xu W, Huang AC, et al. (2018). Feasibility of monitoring advanced melanoma patients using cell-free DNA from plasma. Pigment Cell Melanoma Res, 31: 73-81.
- Valpione S, G Gremel, P Mundra, P Middlehurst, E Galvani, et al. (2018). Plasma total cell-free DNA (cfDNA) is a surrogate biomarker for tumour burden and a prognostic biomarker for survival in metastatic melanoma patients. Eur J Cancer, 88: 1-9.
- Kozak K. (2020). Cell-free DNA BRAF V600E measurements during BRAF inhibitor therapy of metastatic melanoma: long-term analysis. Tumori, 2020: p. 300891619900928.
- Mikoshiba A, Ashida A, Sakaizawa K, Kiniwa Y, Okuyama R. (2020). Detecting copy number alterations of oncogenes in cell-free DNA to monitor treatment response in acral and mucosal melanoma. J Dermatol Sci, 97: 172-178.
- McEvoy AC, Pereira MR, Reid A, Pearce R, Cowell L, et al. (2019). Monitoring melanoma recurrence with circulating tumor DNA: a proof of concept from three case studies. Oncotarget. 10: 113-122.
- Forthun RB, Hovland R, Schuster C, Puntervoll H, Broda HP, et al. (2019). ctDNA detected by ddPCR reveals changes in tumour load in metastatic malignant melanoma treated with bevacizumab. Sci Rep, 9: 17471.
- Zmrzljak UP. (2021). Detection of Somatic Mutations with ddPCR from Liquid Biopsy of Colorectal Cancer Patients. Genes (Basel). 12: 289.
- Palacin-Aliana I. (2024). ddPCR Overcomes the CRISPR-Cas13a-Based Technique for the Detection of the BRAF p.V600E Mutation in Liquid Biopsies. Int J Mol Sci. 25: 10902.
- Kaneko A. (2021). Liquid biopsy-based analysis by ddPCR and CAPP-Seq in melanoma patients. J Dermatol Sci. 102: 158-166.
- Bidshahri R. (2016). Quantitative Detection and Resolution of BRAF V600 Status in Colorectal Cancer Using Droplet Digital PCR and a Novel Wild-Type Negative Assay. J Mol Diagn, 18: 190-204.
- Qin D, Puskas J, Bossler A. (2022). V600E Droplet Digital PCR (ddPCR) cfDNA Troubleshooting. Journal of Molecular Diagnostics, 24: S130-S130.
- Lechner A, Rai A, Rudilla VR, Kuang Y, Paweletz CP, et al. (2024). Atypical Droplet Digital Polymerase Chain Reaction Patterns That Indicate Uncommon but Clinically Actionable EGFR Mutations in Lung Cancer. Arch Pathol Lab Med. 148: 553-558.
© by the Authors & Gavin Publishers. This is an Open Access Journal Article Published Under Attribution-Share Alike CC BY-SA: Creative Commons Attribution-Share Alike 4.0 International License. Read More About Open Access Policy.